Lawrence Berkeley Labs
My latest eBriefing for the New York Academy of Sciences, Growth Networks: Systems Biology Meets Developmental Biology, is now up (the direct link should work only for Academy members; others may get to it through the NYAS page of my website.)
The symposium was very interesting, but, as often happens, it was challenging to present the three talks as a coherent unit. In this case, the overall message (provided by the visionary organizer, Andrea Califano) is that the sweeping and irreversible changes that occur during early development, which are often driven by a relatively few molecular events (perhaps dozens), can provide stringent and useful tests for understanding molecular regulation. This is quite a different way to learn about networks than by gently poking ("perturbing," for example with stress or drugs or RNA interference) a mature animal, in which various molecules are generally cooperating to keep things stable.
The hope is that the overlap between development and systems biology, can have the sort of powerful synergy that have enriched evolutionary and developmental biology in Evo Devo, as popularized by Sean B. Carroll and others. But I suspect the final synthesis will be more of a three way combination, SysEvoDevo.
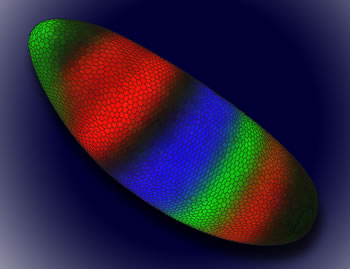
Angela DePace of Harvard, for example, described her nascent efforts to exploit evolutionary comparisons between related species from the fly genus Drosophila, which have been a playground for development (once called embryology) for nearly a century. In the past couple of decades researchers have learned how to modify particular genes so they produce fluorescent molecules of various colors along with their normal protein products. The results have shown in living color how various transcription factors interact to generate that spatial patterns and compartments that ultimately shape the segmented body of the fly. DePace and her former colleagues at Lawrence Berkeley Labs refined the technique to let them measure the quantitative changes in gene expression at thousands of individual cells in the early embryo (see the figure), which let them test the models of gene activity (and the differences between species) in fascinating detail.
Stanislav Shvartsman of Princeton also looked at early Drosophila development, but he showed that the transcription factors alone don't explain everything. Instead, some of the patterning depends on protein phosphorylation, which is a half-century old process that among other things carries signals from a cell's outer membrane to its nucleus, but is rarely considered in development. Antonio Iavarone of Columbia studies the development of the early nervous system in mice from stem cells to differentiated neurons. This is a process that is subverted by brain cancers, which re-activate this cellular program to grow and nourish themselves.
Pulling these three diverse talks together was a bit of a shoe-horning exercise, but they were all fascinating.




No comments:
Post a Comment